Genomicus continues his presentation of the front-loading hypothesis:
___________
Geno: >>In my previous article on the subject of front-loading, I described the front-loading hypothesis and what it proposes. I outlined three testable predictions generated by the front-loading hypothesis. In this article, we’ll see how the front-loading hypothesis can lead us to numerous research questions, and this, in turn, will allow us to establish a better picture about the history of the origin and development of biological complexity.
There are probably dozens of research questions that we can ask as a result of the front-loading hypothesis, so I’ll only cover some of them here.
- How could molecular machines and systems be front-loaded?
An interesting question from a front-loading perspective is how molecular machines and systems (e.g., cilia, etc.) could have been front-loaded. Cilia, for example, usually require a fairly large number of proteins for their assembly and function (IFT particles). While IFT proteins are not required in the cilia of all eukaryotes, they are present in most eukaryotic cilia, and thus, if the cilium was front-loaded, it’s likely that these IFT particles were front-loaded with it.
The basic problem with front-loading a molecular machine is this: if a molecular machine functions through four components, for example, say A, B, C, and D, then how can we arrange it such that, in the future, components A, B, C, and D associate and form a novel molecular machine? At this point, one might be tempted to ask why we can’t just design the molecular machine into the first cells. However, when it comes to many molecular systems, it’s difficult to see how we could plug this molecular system, that is supposed to function in a eukaryotic context, into a prokaryote. For example, could sarcomeres really function in prokaryotes, without drastically re-engineering the sarcomeres such that they can function in prokaryotes? Almost certainly not. Thus, with a number of molecular systems, it seems that they have to be front-loaded, instead of designed into the initial state.
So, how do we front-load molecular machine X, composed of components A, B, C, and D? This research problem is most interesting, actually. If the solution to this problem was solved, it would, at a stroke, offer an explanation for a paradox of sorts: namely, the paradox that while it’s difficult to see how sophisticated molecular machines that display properties of rational design could arise through purely non-teleological mechanisms, often the components of these molecular machine share significant sequence similarity with other molecular systems, suggesting that common descent played a major role in their origin. In other words, molecular systems that are pretty hard to explain through non-teleological mechanisms show signs of having arisen through common descent. The solution to the above problem would solve this paradox.
There are two basic solutions to the problem of front-loading sophisticated molecular systems, but more research is needed so that we can find out exactly how these solutions would work in practice. In theory, however, there’s the “bottom up” approach and the “top down” approach to front-loading molecular systems. In the “bottom up” approach, the original cells would contain the components of the molecular machine we want to front-load, but these components would be carrying out functions not related to the function of the molecular machine. Then, somehow (here’s where we need research), something causes them to associate such that they fit nicely with each other, forming a novel molecular machine.
The “top down” approach proposes that the first cells had a highly complex molecular machine, composed of, say, components A, B, C, D, E, F, G, H, and J. If we want to front-load a molecular machine composed of components A, B, C, and D, then this highly complex molecular machine contains a functional subset of A, B, C, and D. In other words, components E, F, G, H, and J would simply have to be deleted from the highly complex molecular machine, resulting in a molecular machine composed of A, B, C, and D. This model is actually testable. Under this model, we would tentatively predict that a homologous system of a molecular machine will be more complex if it is more ancient than the molecular machine.
The nice thing about the “top down” approach is that we might just have an actual example of this in the biological world.
Consider the bacterial flagellum. It shares homology with the type III secretion system (TTSS), a molecular nanomachine that is involved in interactions between bacteria and eukaryotes (the TTSS exports proteins through the inner and outer membrane of the bacterium into the host cell). Might the TTSS have been front-loaded? This is an interesting possibility since it doesn’t seem all that difficult for bacteria to evolve export systems from the flagellum. Thus, the bacterium Buchnera aphidicola, an endosymbiont of the pea aphid, is nonmotile although its genome does have a cluster of flagellar genes. Research by Maezawa et al. (2006) indicates that these flagellar genes are transcribed and translated, and form a hook-basal-body complex (HBB) on the cell surface of Buchnera. In other words, at least two protein export systems have evolved from the flagellum independently, highlighting the possibility that, through the “top down” approach, molecular machines may be fairly easy to front-load.
Now, before someone mentions that the TTSS is involved in virulence, I’d like to point out that, if the TTSS was front-loaded, promoting virulence was probably not its intended function. Instead, something like beneficial symbiosis was probably its original function (in a front-loading context), and vestiges of this original state can be found in the fact that the TTSS mediates beneficial interactions between bacteria and legumes.
- Has cytosine deamination played an important role in the origin of metazoan genes?
Cytosine deamination, wherein the DNA base cytosine changes to thymine, is one the most common types of mutations. Poole et al. argued that no engineer would have used cytosine as a base in DNA precisely because it is prone to deamination. However, Mike Gene noted that cytosine deamination actually changes a pool of various amino acids to a pool composed of almost entirely strongly hydrophobic amino acids (called the “Increasing Hydrophobicity Effect,” or IHE). Given that hydrophobic residues are known to be play key roles in protein folding, this interesting discovery suggests, from a front-loading perspective, that cytosine may have been chosen by an engineer(s) because it is prone to deamination. This hypothesis predicts, then, that cytosine deamination has played a fairly important role in the origin of metazoan genes. This hypothesis can be tested by changing a metazoan gene’s sequence into its hypothesized original state before it was hit by cytosine deamination, and then BLASTing this new sequence to see if we get any significant prokaryotic homologs. Such a find would strengthen this hypothesis.
- Can the IHE be turned on and off by the cell?
Cytosine deamination is corrected by an enzyme called uracil glycosylase, and it can be increased by the enzyme cytidine deaminase. This brings up the possibility that the cell itself can control such molecular machinery, and thereby controlling the “increasing hydrophobicity effect.” This would, in theory, enable the cell to rapidly evolve certain proteins when it receives a certain stimulus. Thus, if the front-loading intelligence had designed such a system into the cell, we would predict that uracil glycosylase and cytidine deaminase was present in the very first cells, and, as a consequence, we would expect these enzymes (or enzymes carrying out the same function) to be found in the deepest-branching life forms, and to be nearly as widespread as ATP synthases.
- What was front-loaded?
This particular question is quite difficult to resolve, although the front-loading hypothesis does posit that metazoan life forms were front-loaded, and almost certainly animals and plants were front-loaded. Beyond this, it is difficult to say what was front-loaded. Were all animal and plant phyla front-loaded or only certain ones? Were various classes also front-loaded? How hard would it be to front-load a specific animal class, for example? Was the bacterial flagellum front-loaded, or was it designed into the first cells (note: I opt for the latter idea, that the flagellum was designed into the first cells)?
- What were the first cells?
Under the front-loading hypothesis, what did the first cells look like? Were they bacteria, archaea, or neither? Or perhaps the first cells did not belong to the same phyla and domains. What kind of genomic information did they contain? Cyanobacteria make an interesting candidate for the first cells, for several reasons. Firstly, they are deep-branching, which means they could have been there at the dawn of life on earth (in fact, cyanobacteria fossils are some of the oldest fossils out there). Secondly, they play a key role in the biosphere, suggesting that they could have easily been used to terra-form the earth such that it would be more hospitable to future, complex life forms. Thirdly, cyanobacteria often come together to form colonies, which could, I suppose, be useful in a front-loading context.
It is my opinion that the initial cells were composed of different types of bacteria, not belonging to any one group.
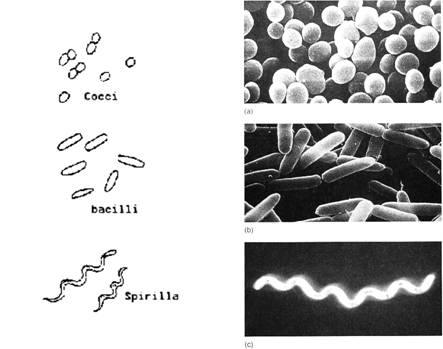
Figure. A figure of different types of bacteria. Image source: here.
Conclusion
In this article, I covered five different research questions we can ask from a front-loading angle. Answering these questions will be useful for a further understanding of how biological complexity arose. Also, many of these research questions offer testable predictions which is a good thing. If, for example, it was found that cytosine deamination has played an important role in the origin of metazoan genes, this would be a good reason for a front-loading designer to include cytosine as a base in DNA, countering the objection of Pool et al., and offering evidence for the front-loading hypothesis.
About me
Over the years, I have become quite interested in the discussion over biological origins, and I think there is “something solid” behind the idea that teleology has played a role in the history of life on earth. When I’m not doing multiple sequence alignments, I’m thinking about ID and writing articles on the subject, which can be found on my website, The Genome’s Tale.
I am grateful to UD member kairosfocus for providing me with this opportunity to make a guest post on UD. Many thanks to kairosfocus.
Also see The Design Matrix, by Mike Gene.
References
- Kazuki Maezawa, Shuji Shigenobu, Hisaaki Taniguchi, Takeo Kubo, Shin-ichi Aizawa, and Mizue Morioka. Hundreds of Flagellar Basal Bodies Cover the Cell Surface of the Endosymbiotic Bacterium Buchnera aphidicola sp. Strain APS. J. Bacteriol., 2006, 188(18): 6539–6543.
- Poole A, Penny D, Sjöberg BM. Confounded cytosine! Tinkering and the evolution of DNA. Nat Rev Mol Cell Biol., 2001; 2(2):147-51.>>
_________________
Stay tuned for more . . . END