Let’s read the Nature abstract:
Nature (2019) Article | Published: 15 May 2019
Total synthesis of Escherichia coli with a recoded genome
Julius Fredens, Kaihang Wang, Daniel de la Torre, Louise F. H. Funke, Wesley E. Robertson, Yonka Christova, Tiongsun Chia, Wolfgang H. Schmied, Daniel L. Dunkelmann, Václav Beránek, Chayasith Uttamapinant, Andres Gonzalez Llamazares, Thomas S. Elliott & Jason W. Chin
Abstract
Nature uses 64 codons to encode the synthesis of proteins from the genome, and chooses 1 sense codon—out of up to 6 synonyms—to encode each amino acid. Synonymous codon choice has diverse and important roles, and many synonymous substitutions are detrimental. Here we demonstrate that the number of codons used to encode the canonical amino acids can be reduced, through the genome-wide substitution of target codons by defined synonyms. We create a variant of Escherichia coli with a four-megabase synthetic genome through a high-fidelity convergent total synthesis. Our synthetic genome implements a defined recoding and refactoring scheme—with simple corrections at just seven positions—to replace every known occurrence of two sense codons and a stop codon in the genome. Thus, we recode 18,214 codons to create an organism with a 61-codon genome; this organism uses 59 codons to encode the 20 amino acids, and enables the deletion of a previously essential transfer RNA. [Cited, per fair use doctrine for academic, non commercial purposes.]
Let us refresh memory on the genetic code:
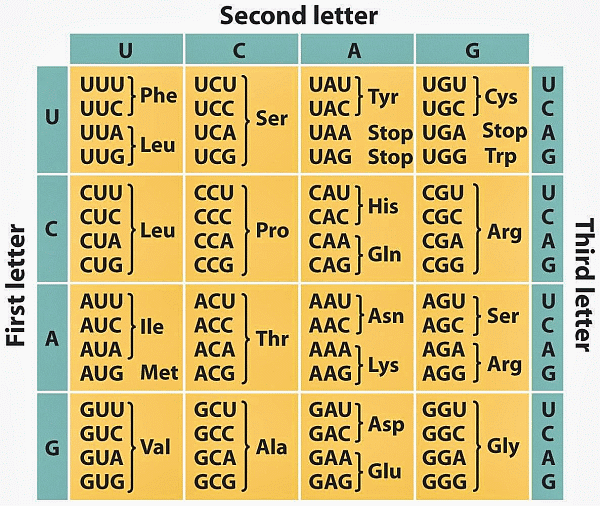
And on the DNA:
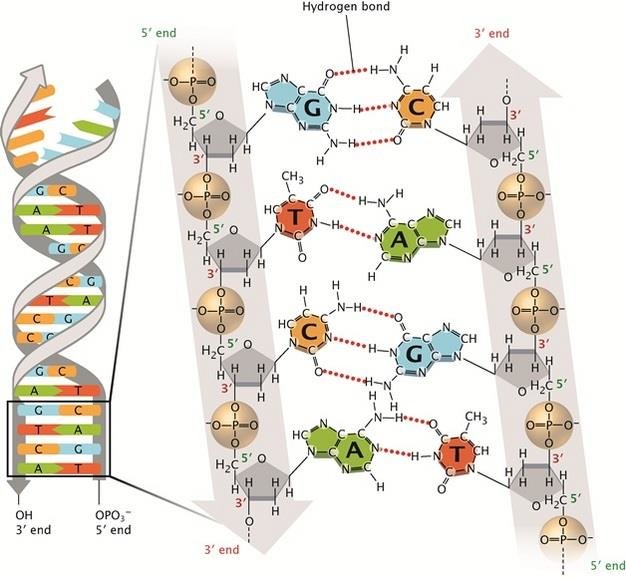
Then also, protein synthesis:
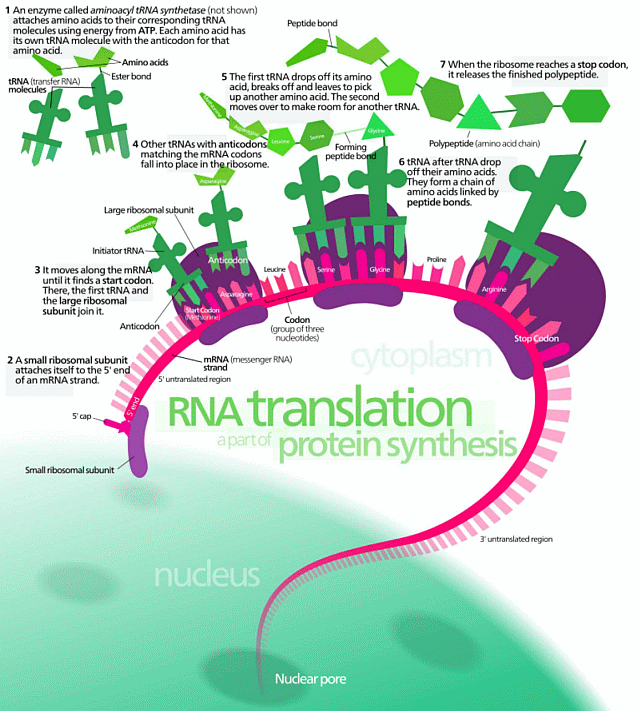
Phys dot org gives some context:
A team of researchers at Cambridge University has replaced the genes of E. coli bacteria with genomes they synthesized in the lab. In their paper published in the journal Nature, the group describes replacing the genome and removing redundant genetic codes [–> three letter 4-state elements have 64 possibilities but only 20 are needed for typical protein AA’s, AUG codes for an AA and serves as START, there are three STOP codons] . . . . In this new effort, the researchers had two goals: The first was to synthesize the genome of an E. coli bacterium in their lab—all four million letters of it. The second was to find out what would happen to such a specimen if some of its DNA redundancies were removed . . . .
The researchers report that it took longer for the special bacterial specimen to grow, but other than that, it behaved just like unedited specimens. They suggest that in future efforts, it might be possible to replace the redundancies they removed with other sequences to create bacteria with special abilities, such as making new types of biopolymers not found in nature.
In short, they confirmed that the choice of “synonym” has a regulatory effect.
Where are we today, then?
First, we have definitive demonstration of the intelligent design of a genome. Yes, they obviously have not created a de novo cell body (a much more difficult task), but we see that intelligent design of life here definitively passes the Newton test of observed actual cause. Further, we see that DNA functions as an information system in the cell, supporting the significance of this conceptual representation, based on Yockey’s work:
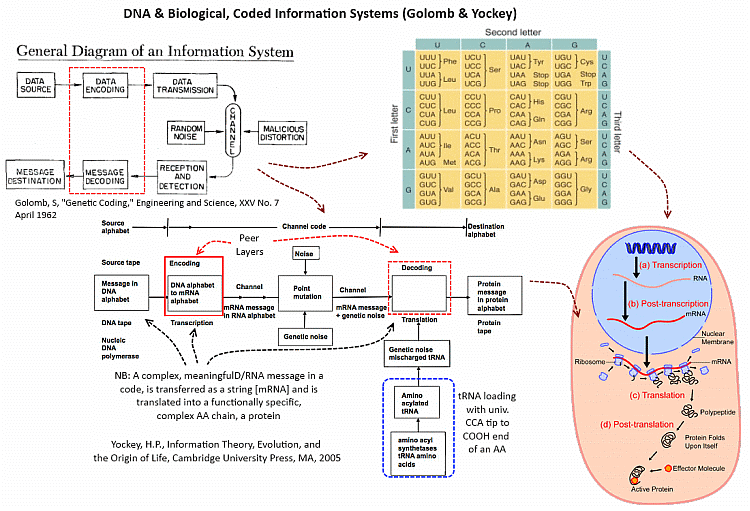
I add: Let’s zoom in on Yockey’s contribution, on the code-communication system as applied to protein synthesis, which underscores the linguistic nature of what is involved:
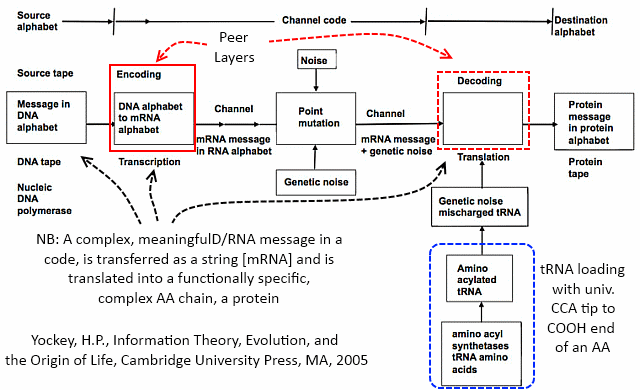
Where, Crick understood this from the beginning in 1953, witness p. 5 of his letter to his son Michael, March 19, 1953:

At this stage, we definitively know that using nanotech molecular biology and linked computational techniques it is feasible to construct a genome based on intelligently directed configuration. AKA, design.
Therefore, intelligent design, as of right not sufferance, sits at the table for study on origin of life and of body plans.
Where, we separately know on configuration space search challenge, that it is maximally implausible to construct in excess of 500 – 1,000 bits of functionally specific complex organisation and/or associated information. As a reminder:
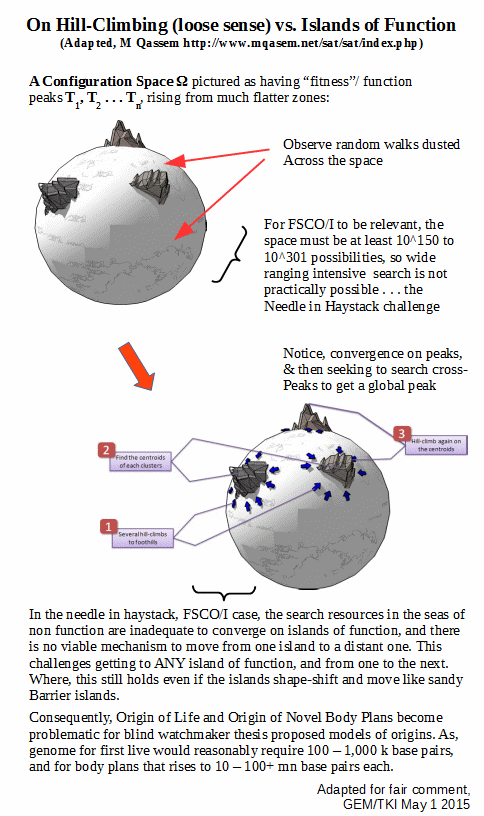
We are now in a different ball game completely: Intelligent Design of life is demonstrated to be feasible and actual in the here and now, as of this investigation. Therefore, as of right, it is a serious candidate to explain what we see in the world of life; especially as regards origin of cell based life and origin of main body plans.
Going forward, we are now a full-fledged independent school of thought. END
PS: James Tour on the Mystery of Life’s Origin, challenging the usual OoL claims, focus from c. 8:30 on:
PPS: It seems we need to understand that there are such things as DNA Synthesisers. Here, is a sample, the “Dr Oligo”:
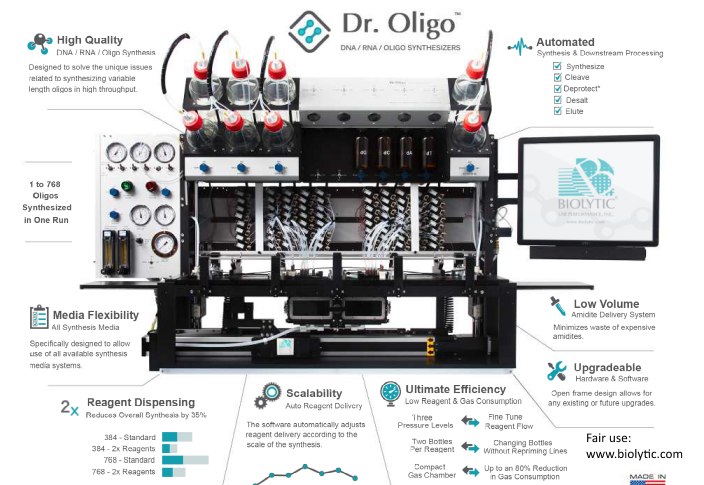

Biocyclopedia lays out the architecture:
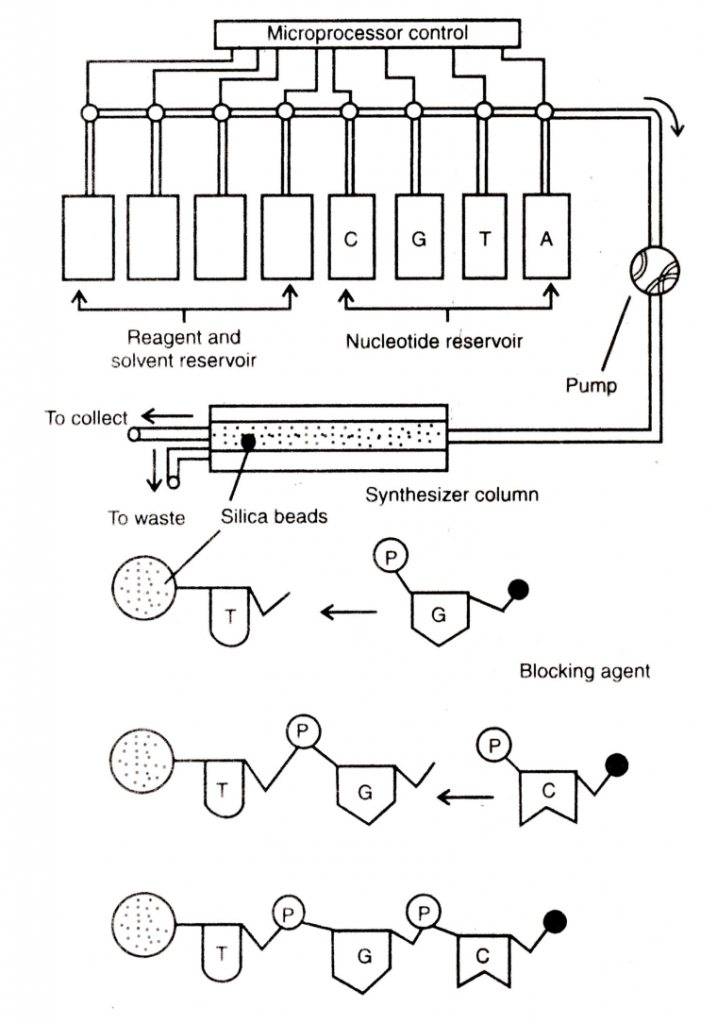
Clipping the explanation:
Recently, fully automated commercial instrument called automated polynucleotide synthesizer or gene machine is available in market which synthesizes predetermined polynucleotide sequence. Therefore, the genes can be synthesized rapidly and in high amount. For example, a gene for tRNA can be synthesized within a few days through gene machine. It automatically synthesizes the short segments of single stranded DNA under the control of microprocessor. The working principle of a gene machine includes (i) development of insoluble silica based support in the form of beads which provides support for solid phase synthesis of DNA chain, and (ii) development of stable deoxyribonucleoside phosphoramidites as synthons which are stable to oxidation and hydrolysis, and ideal for DNA synthesis.
The mechanism of a gene machine is shown in Fig. 2.14 [–> above]. Four separate reservoirs containing nucleotides (A,T,C and G) are connected with a tube to a cylinder (synthesizer column) packed with small silica beads. These beads provide support for assembly of DNA molecules. Reservoirs for reagent and solvent are also attached. The whole procedure of adding or removing the chemicals from the reagent reservoir in time is controlled by microcomputer control system i.e. microprocessor . . . .The desired sequence is entered on a key board and the microprocessor automatically opens the valve of nucleotide reservoir, and chemical and solvent reservoir. In the gene machine the nucleotides are added into a polynucleotide chain at the rate of two nucleotides per hour. By feeding the instructions of human insulin gene in gene machine, human insulin has been synthesized.
As in, molecular nanotech lab in action.
PPPS: As objectors have raised the claimed logical, inductive inference that designing intelligences are embodied (which we can safely hold, implicitly “lives” in the context of the presumed, evolutionary materialistic account of origins — of cosmos, matter, life, body plans, man, brains and minds), I first link a discussion of how this undermines rationality, by Craig:
I also put on the table the Smith, two-tier supervisory controller bio-cybernetic model, as a context to discuss embodiment, intelligence and computational substrates, first in simplified form:
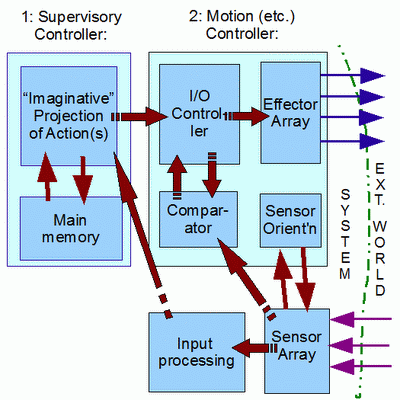
Then, in more full detail:
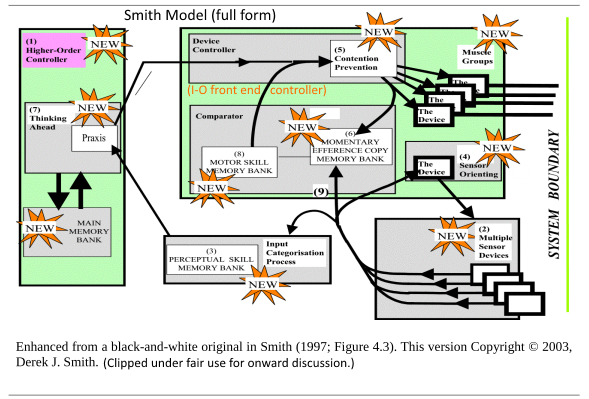
This then leads to the gap between computation on a substrate and rational contemplation. That is, Reppert’s point holds:
. . . let us suppose that brain state A [–> notice, state of a wetware, electrochemically operated computational substrate], which is token identical to the thought that all men are mortal, and brain state B, which is token identical to the thought that Socrates is a man, together cause the belief [–> concious, perceptual state or disposition] that Socrates is mortal. It isn’t enough for rational inference that these events be those beliefs, it is also necessary that the causal transaction be in virtue of the content of those thoughts . . . [But] if naturalism is true, then the propositional content is irrelevant to the causal transaction that produces the conclusion, and [so] we do not have a case of rational inference. In rational inference, as Lewis puts it, one thought causes another thought not by being, but by being seen to be, the ground for it. But causal transactions in the brain occur in virtue of the brain’s being in a particular type of state that is relevant to physical causal transactions.