I do not take a decisive position on the extent to which taxonomic groups are related by descent. As such, my current position is one of skeptical agnosticism. While I view neo-Darwinian evolution as devoid of scientific traction, I think that common descent is still — at the very least — defensible, and on this issue I am quite happy to be persuaded one way or the other. Now, there is no doubt that universal common ancestry faces a lot of problems: such as the widely divergent early vertebrate embryonic stages and the non-congruence between homology and developmental pathways. But there are also some meritorious arguments for more modest levels of common ancestry — such as at the primate level. This is not to say that I think such arguments are iron-clad — far from it. Indeed, in my various postings on Evolution News & Views and Uncommon Descent, I have sought to demonstrate why the arguments for common descent are NOT conclusive.
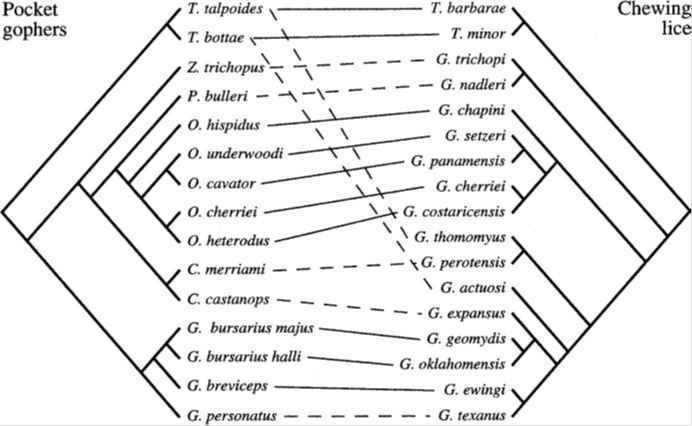
One argument which I have, for a while, regarded as among the stronger of the arguments for common ancestry is the phenomenon of host and parasite cospeciation — which yields correlating phylogenetic trees. For example, take a look at the following diagram excerpted from Legendre et al. (2001):
The diagram (called a “tanglegram”) represents the parallel phylogenies yielded by the rodents “pocket gophers” and their lice parasites. The lines in the middle connect each species of pocket gopher to their respective parasites. As can be seen, with rare exception, the alignment of the trees is remarkably good. And occasionally one observes a host-switch event. Similar studies have been conducted for quite a number of other groups such as parasites affecting song birds and primates, as well as figs and fig wasps.
Now, the sheer number of potential phylogenies for those groups renders appeals to chance void. And it is widely thought that common ancestry (and cospeciation) provides the only conceptual framework for understanding these parallel phylogenies.
But does it?
In 2000, an interesting paper was published by Sharp et al. This research team sought to determine the divergence times of primate lentivirus (a genus of retroviruses which includes HIV and SIV). They found that, while the lentivirus phylogeny closely paralleled the primate phylogeny, the divergence times were completely different. The authors reported,
In the absence of a fossil record, two main approaches have been used in attempts to establish time-scales for viral evolution. For rapidly evolving viruses, it is possible to calibrate their rate of evolution by comparison of viruses isolated at different time points, and then to use this molecular clock to estimate the dates of divergences within the evolutionary tree. This procedure has been used, for example, with influenza A viruses[13].The other approach has been to find instances where viruses appear to have been evolving in a host-dependent manner, i.e. without transmission between different species of host. In such cases, the divergence times of hosts also pertain to their viruses. It has been suggested that this method can be applied to herpesviruses [14].
Both techniques have been adopted to estimate the time-scale of primate lentivirus evolution. Some of the earliest attempts, in 1988, yielded dramatically discordant results. One attempt to use a molecular clock estimated that the common ancestor of HIV-1 and HIV-2 existed as recently as 1951 [15]. At the same time, after comparison of HIV-1 with an SIV from an African green monkey, others suggested that these viruses may have diverged `gradually in concert with the evolution of primates’ [16], and that it was possible that a common ancestor of these viruses infected the common ancestor of apes and Old World monkeys [17]; that ancestor is thought to have lived at least 25 million years ago. Since HIV-1, HIV-2 and SIVAGM each belong to different major lineages of primate lentivirus (Figure 1), these two estimates pertain to the same common ancestor of all the primate lentiviruses. Thus the two different approaches led to estimates with a discrepancy of about six orders of magnitude! [emphasis added]
How is one to account for this discrepancy? One suggestion that has been put forward is the idea that our modern molecular dating methodologies are extremely faulty. But it’s not likely that this significant discrepancy could be overcome simply by enhanced dating techniques.
In 2002, in the Journal of Systematic Biology, Charleston and Robertson published a paper entitled “Preferential Host Switching by Primate Lentiviruses Can Account for Phylogenetic Similarity with the Primate Phylogeny.” In the paper, they write,
If codivergence is insuffcient to explain the similarity between the virus and host phylogenies, could any other process generate such coincidental results? We present here an alternative hypothesis of preferential host switching among primate host lineages, with host switching more likely to be successful between more closely related hosts than between more distantly related hosts. This hypothesis is used in a simulation study in which an artifcial virus phylogeny is “grown” on the primate host phylogeny by a process of divergence (not codivergence) and host switching. We demonstrate how this simple principle can lead to artificially similar looking host and virus phylogenies, which can erroneously suggest codivergence where none occurred. [emphasis added]
They thus present the view that cross-species transmission occurs preferentially between genetically similar species, thus producing the illusion of host/parasite cospeciation. They further suggest that “There may well be other cases of host/parasite coevolution in which this phenomenon occurs.”
This phenomenon adds but one further example to the growing trend: Many of the purported cases of “evidence for common ancestry” have plausible alternative explanations — a fact which sits comfortably within the context of neo-Darwinism’s increasingly dubious foundations as a causal explanation for what we find in biology. No doubt further research will reveal this to be only the peak of the ice berg.