Extreme genome repair, and remarkable morphogenesis by self-assembly point to design
https://reasonandscience.catsboard.com/t2061p275-my-articles#9626
Extreme Genome Repair (2009): If its naming had followed, rather than preceded, molecular analyses of its DNA, the extremophile bacterium Deinococcus radiodurans might have been called Lazarus. After shattering of its 3.2 Mb genome into 20–30 kb pieces by desiccation or a high dose of ionizing radiation, D. radioduransmiraculously reassembles its genome such that only 3 hr later fully reconstituted nonrearranged chromosomes are present, and the cells carry on, alive as normal 1
T. Devitt (2014): John R. Battista, a professor of biological sciences at Louisiana State University, showed that E. coli could evolve to resist ionizing radiation by exposing cultures of the bacterium to the highly radioactive isotope cobalt-60. “We blasted the cultures until 99 percent of the bacteria were dead. Then we’d grow up the survivors and blast them again. We did that twenty times,” explains Cox. The result were E. coli capable of enduring as much as four orders of magnitude more ionizing radiation, making them similar to Deinococcus radiodurans, a desert-dwelling bacterium found in the 1950s to be remarkably resistant to radiation. That bacterium is capable of surviving more than one thousand times the radiation dose that would kill a human. 2
Simple bacteria can restart their ‘outboard motor’ by hotwiring their own genes (2015):
Unable to move and facing starvation, the bacteria evolve a replacement flagellum – a rotating tail-like structure that acts like an outboard motor – by patching together a new genetic switch with borrowed parts. When an organism suffers a life-threatening mutation, it can rapidly rewire its genes. The remarkable speed with which old genes take on new tasks suggests that life has unexpected levels of genetic flexibility. In theory, the bacteria should have starved to death and effectively gone extinct. Yet over the course of a weekend, they managed to patch themselves back together with borrowed genes.” Scientists made the discovery by accident while researching ways to use naturally occurring bacteria to improve the yield of crops. A microbe was engineered so that it could not make its ‘propeller-like’ flagellum and forage for food. However, when a researcher accidentally left the immotile strain out on a lab bench, the team discovered the bacteria had evolved over just a few days. The new variety of bacteria had resurrected their flagella in the process.
Remarkably, this happened because the mutants had rewired a cellular switch, which normally controls nitrogen levels in the cell, to activate the flagellum. This rescued these bacteria, which faced certain death if they didn’t move to new food sources. The bacteria being studied, Pseudomonas fluorescens, are among a group of bacteria scientists are researching for use in agriculture, as a kind of ‘plant probiotic’. These could help crops grow or fight off diseases, leading to higher yields. However, a key problem is that the bacteria lack resilience, as their positive effects can stop working after only a short period of time. Dr Jackson, a microbiologist at Reading, said: “Plant probiotics could make crops grow more reliably in the future, helping to feed the world’s growing population. This new study shows that these bacteria are more resilient than previously thought, as they show a remarkable capacity to overcome catastrophic changes and find a way to survive. “This gives us crucial insights into how bacteria could survive and change, and the challenge now is to see if this occurs in their natural soil and plant environment.” 3
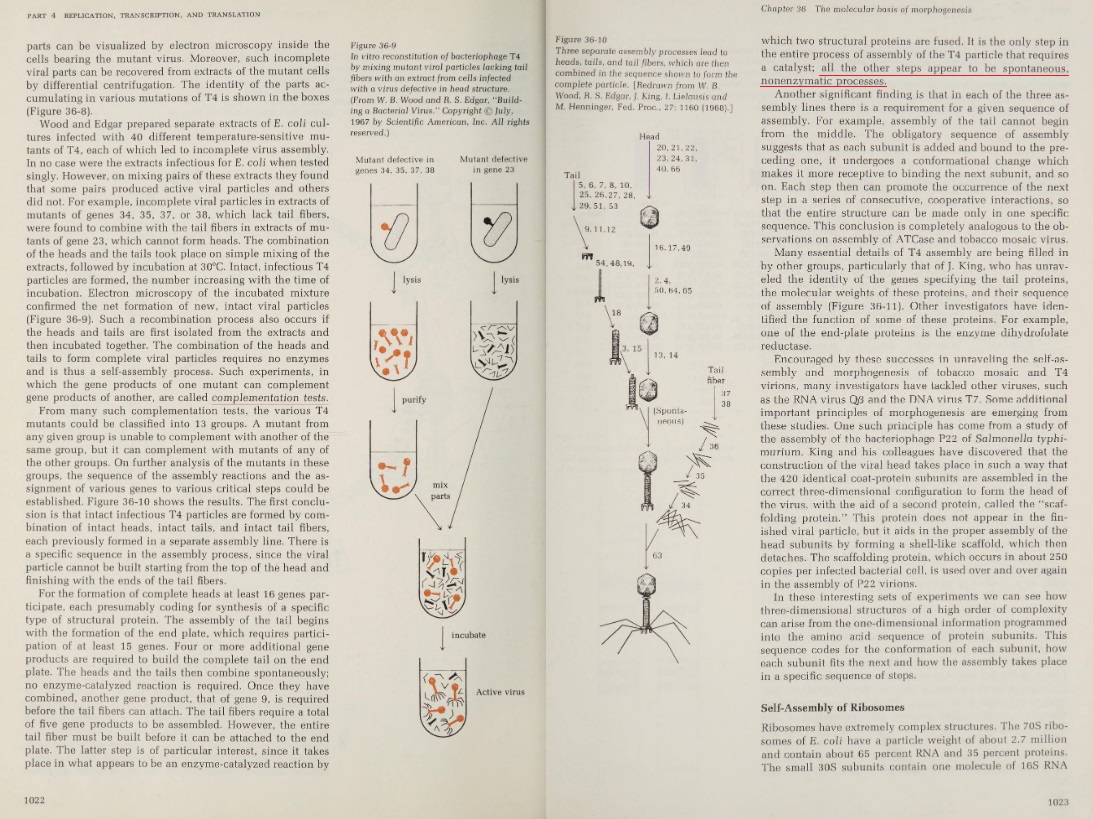
K. Eric Drexler: Engines of Creation 2.0 ( 2006): The T4 phage, acts like a spring-loaded syringe and looks like something out of an industrial parts catalog. It can stick to a bacterium, punch a hole, and inject viral DNA (yes, even bacteria suffer infections). Like a conqueror seizing factories to build more tanks, this DNA then directs the cell’s machines to build more viral DNA and syringes. Like all organisms, these viruses exist because they are fairly stable and are good at getting copies of themselves made. Whether in cells or not, nanomachines obey the universal laws of nature. Ordinary chemical bonds hold their atoms together, and ordinary chemical reactions (guided by other nanomachines) assemble them. Protein molecules can even join to form machines without special help, driven only by thermal agitation and chemical forces. By mixing viral proteins (and the DNA they serve) in a test tube, molecular biologists have assembled working T4 viruses. The machinery of the T4 phage, for example, self-assembles from solution, apparently aided by a single enzyme.…self-assembling structures (…For a description of molecular self-assembly, including that of the T4 phage and the ribosome, see Chapter 36 of Lehninger’s Biochemistry)
This ability is surprising: imagine putting automotive parts in a large box, shaking it, and finding an assembled car when you look inside! Yet the T4 virus is but one of many self-assembling structures. (M. YANAGIDA, 1984: The virus particle contains more than 3,000 protein subunits of some 30 polypeptide species !!) Molecular biologists have taken the machinery of the ribosome apart into over fifty separate protein and RNA molecules, and then combined them in test tubes to form working ribosomes again. To see how this happens, imagine different T4 protein chains floating around in water. Each kind folds up to form a lump with distinctive bumps and hollows, covered by distinctive patterns of oiliness, wetness, and electric charge. Picture them wandering and tumbling, jostled by the thermal vibrations of the surrounding water molecules. From time to time two bounce together, then bounce apart. Sometimes, though, two bounce together and fit, bumps in hollows, with sticky patches matching; they then pull together and stick. In this way protein adds to protein to make sections of the virus, and sections assemble to form the whole. 4
E. V. Koonin, the logic of chance, page 376: Breaking the evolution of the translation system into incremental steps, each associated with a biologically plausible selective advantage is extremely difficult even within a speculative scheme let alone experimentally. Speaking of ribosomes, they are so well structured that when broken down into their component parts by chemical catalysts (into long molecular fragments and more than fifty different proteins) they reform into a functioning ribosome as soon as the divisive chemical forces have been removed, independent of any enzymes or assembly machinery – and carry on working. Design some machinery that behaves like this and I personally will build a temple to your name! 5
AlphaFold (2020) What a protein does largely depends on its unique 3D structure. Figuring out what shapes proteins fold into is known as the “protein folding problem”, and has stood as a grand challenge in biology for the past 50 years. In a major scientific advance, the latest version of our AI system AlphaFold has been recognized as a solution to this grand challenge by the organizers of the biennial Critical Assessment of protein Structure Prediction (CASP). 6
Comment: Imagine the engineering effort that it would take for protein engineers to produce nanomachines that would need no nano arms and nano hands to assemble complex nanomachines, but design parts that would be able to assemble on their own just by shaking them, like motors, bearings, and moving parts coming together randomly, and then self-assemble into a fully operational nano-machine. The engineers would need to know the single individual forces and how they would interact with the forces from the other parts. The problem becomes even more apparent when we consider that one of the forces that influence proteins is for example Van der Waals forces which operate based on quantum mechanical principles. R. W. Newberry (2019): The dominant contributors to protein folding include the hydrophobic effect and conventional hydrogen bonding, along with Coulombic interactions and van der Waals interactions. Human technology and advance is far from being able to design this. What a feat would THAT be!
1. Rodrigo S. Galhardo: Extreme Genome Repair 2012 Apr 4.
2. T. Devitt: In the lab, scientists coax E. coli to resist radiation damage March 17, 2014
3. WEEKEND EVOLUTION: BACTERIA ‘HOTWIRE THEIR GENES’ TO FIX A FAULTY MOTOR 26 February 2015
4. K. Eric Drexler: Engines of Creation 2.0 ( 2006)
5. E. V. Koonin, The logic of chance (2012), page 376
6. AlphaFold: a solution to a 50-year-old grand challenge in biology November 30, 2020