As the genome responds to traumatic experiences:
A piece of “junk DNA” could be the key to extinguishing fear-related memories for people struggling with post-traumatic stress disorder (PTSD) and phobia, according to a study from The University of Queensland.
An international research project, led by the Queensland Brain Institute’s Associate Professor Timothy Bredy, discovered the new gene while investigating how the genome responds to traumatic experiences.
“Until recently, scientists thought the majority of our genes were made up of junk DNA, which essentially didn’t do anything.” Dr. Bredy said.
“But when researchers began to explore these regions, they realized that most of the genome is active and transcribed.”
University of Queensland, “‘Junk DNA’ could be key to controlling fear” at Phys.org (March 22, 2022)
Okay, why, until recently, did researchers think that “the majority of our genes were made up of junk DNA, which essentially didn’t do anything”?
Because that vast sunken library of dead information (sheer randomness and waste) was a slam dunk for Darwinism, as politically powerful theistic evolutionist Francis Collins was quick to point out in The Language of God. (2007). To say nothing of atheist cultural icon Richard Dawkins here, Darwinian evolutionary biologist Jerry Coyne (here), and unidirectional skeptic Michael Shermer (here). Notice how that history is quietly being erased. Otherwise, it would be necessary to acknowledge that what many regarded as a correct prediction from Darwinism is not true.
So now what about fear?
“Our findings suggest that long non-coding RNAs provide a bridge, linking dynamic environmental signals with the mechanisms that control the way our brains respond to fear,” Dr. Bredy said.
“With this new understanding of gene activity, we can now work towards developing tools to selectively target long non-coding RNAs in the brain that directly modify memory, and hopefully, develop a new therapy for PTSD and phobia.”
University of Queensland, “‘Junk DNA’ could be key to controlling fear” at Phys.org (March 22, 2022)
Here’s the proposed mechanism:
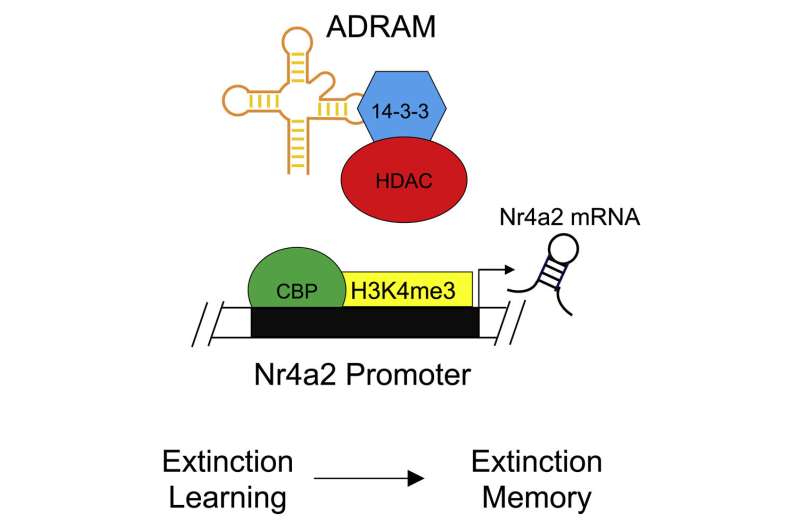
Certainly worth pursuing in terms of addressing PTSD and phobias, as the authors note.
The paper is open access.
You may also wish to read: A new, useful, description for (former) junk DNA… ? “the large proportion of our genome that does not instruct our cells to form proteins” The phrase is a bit longish, of course, but concision is usually a product of usage. It’s better than “non-coding DNA” because it’s more specific and limited as a privative. That is, there is a specific thing that that vast mass of DNA does not do. The longish phrase does not come with the implication that it doesn’t do anything.